This last week we learned of two Manhattan high school students who, in a science project, gave us a glimpse of the future. They asked a simple question: “When you buy fish, do you get the kind of fish you think you are buying/” Anyone who has bought fish at the market or ordered fish at a sushi restaurant knows the problem in the store, one fish looks pretty much like another. In particular, some of the more trendy, tasty and expensive kinds of fish look a lot like cheaper substitutes. White tuna, for example, is difficult to distinguish in the store from farm-raised tilapia, although you pay a lot more for it.
How is a person to know which is which? Animals and plants don’t come with easy-to-read labels telling you to which species an individual belongs. One solution to this dilemma would be to take the specimen to a professional fish taxonomist, who will evaluate fin shape, scales, and many other characters to correctly identify the fish. However, there are not all that many fish taxonomists, and certainly not that many to be found in sushi restaurants.
The bar code
Enter Dr. Paul Hebert of the University of Guelph in Ontario, Canada, with a deceptively simple suggestion, which is that organisms do in fact come with easy-to-read labels. Certain genes vary little among individuals of a species, while different species have different versionswhy not let these genes serve as ID tags? DNA sequencing machines read off the order of nucleotides in batches of about 650 bases at a time, so he proposed examining the first 648 nucleotides of a gene called cytochrome c oxidase subunit 1 (CO1). Why this particular gene? Because CO1 differences between individuals within a species are surprisingly smalljust 2% of individuals differ at all along the 648 stretch of nucleotides. This within-species uniformity is unusualin a typical gene, many differences would be found between members of a species, so many as to overlap with individuals of closely related species. Not so for CO1. Perhaps because cytochrome oxidase play such a critical role in oxidative metabolism, any changes within a species are rare, and when they do occur, spread rapidly through all members of the species. However, after two species have diverged, rare changes that occur in one species do not spread to the other, and so the two species accumulate defining differences.
How well does Hebert’s suggestion work in practice? As a practical test, he set out to compare the first 648 nucleotides of the CO1 gene in mitochondrial DNA obtained from museum specimens of birds. He characterized 341 of the 709 known species of North American birds, and in every instance found a unique sequence characteristic of the species. You can see below the photographs the sequences Hebert found for two marsh wrens. Eastern and Western Marsh Wrens differ in 21 places, indicated by the lines between the two colored “bar codes.” Just as Hebert proposed, the CO1 sequences are easy-to-read labels that clearly identify an individual as being one kind of marsh wren, and not the other.
Dr. Hebert calls his approach “DNA bar coding” by analogy with the bar codes on supermarket items.
Of course, not all organisms are birds. How well does CO1 work as a bar code for other kinds of organisms? So far, it seems to work well for animals, but not as well for plants. Plant biologists have already begun to examine instead two genes found on chloroplast DNA, which seems better suited to distinguishing between plant species. While the central idea of bar coding is the use of a standard gene as a reference, the use of different genes for different major groups presents no major problemfew taxonomists would confuse a bird with a plant.
Using Bar Codes in Taxonomy
At first glance, you might think bar coding will put the world’s taxonomists (people that identify organisms by comparing their bodies and genes) out of business. Not really. While DNA bar codes based on the CO1 gene appear to provide a useful ID tag for birds and other animal species, it is important to be very clear about the nature of the groups the bar codes identify. Any reproductively isolated line of descent can, and indeed would be expected to, develop a unique CO1 bar code. That does not mean the group is a separate species. For example, the aboriginal peoples of Australia have been reproductively isolated for many centuries, but if they were to have developed a unique CO1 bar code, that would not make them a separate species. It would imply no more and no less than genetic isolation.
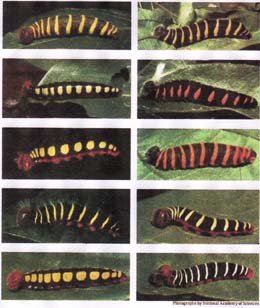
This important distinction can be clearly seen in a 2004 study of Costa Rica butterflies. The skipper butterfly Astraptes fulgerator was first described in 1775 and ranges from Texas to Argentina. Over the course of 25 years of study, University of Pennsylvania ecologist Daniel Janzen has raised some 2,500 caterpillars of the Astraptes skipper, which as you can see in the photograph come in ten versions, each of which feeds on a different plant but all of which give rise to adults that look identical to one another and are all considered members of the same species.
Hearing of the bar code technology being developed by Canadian geneticist Paul Hubert, Janzen tore off one leg from each of his numerous preserved adult skipper butterflies and sent the severed limbs to Dr. Hebert for analysis. Using the same CO1 bar code analysis that had proven so powerful in identifying bird species, Herbert found that Janzen’s skipper collection fell neatly into 10 separate bar code clusters. All members of a cluster had nearly the same bar code, different from the other nine clusters. Importantly, the clusters matched the caterpillar groups! Each type of caterpillar had its own CO1 signature.
The researchers concluded that in Costa Rica there is not one but ten kinds of skipper, each with a strikingly different caterpillar. Janzen speculates that the groups diverged perhaps 4 million years ago, based on the amount of CO1 difference seen, each group specializing on a different caterpillar food plant. Is this a valid way to identify species? No. While genetically unique, there is no evidence these ten kinds of skipper are different enough to be said to be different species. You get the point: the fact that each different species has a unique bar code does not imply that each unique bar code represents a different species.
Bar code forensics
Now, let us return to the tale of the mislabeled fish. Our two intrepid high school sleuths, Kate Stoekle and Louisa Strauss, invested $300 to purchase 60 samples of sea food from four sushi restaurants and ten grocery stores in Manhattan, cut off small portions and sent them to the University of Guelph in Ontario, Canada. There Herbert’s Barcode of Life Database researchers compared the teenagers’ samples with the global library of 30,562 bar codes representing some 5,500 fish species.
Perhaps the results will not surprise you: 2 of the 4 restaurants and 6 of the 10 grocery stores had sold mislabeled fish — one in four of the fish samples were not what they claimed to be! In addition to tilapia sold as white tuna, and smelt sold as flying fish roe, seven of the nine samples that were sold as red snapper were mislabeled as anything from Atlantic cod to Acadian redfish (an endangered species).
The great potential of bar coding is that it allows anyone to correctly identify an unknown specimen in a direct and unambiguous fashion. The testing protocol is very easy, and has been simplified to the point that the high school students could have done it in their kitchen without sending samples off to a place like Guelph. There is even talk of developing simple hand held devices that would allow someone to check a fish right in the restaurant to see if it is the kind of fish claimed on the menu.
And our high school Sherlocks future scientists? Nope. Both are starting college classes this week at Johns Hopkins, one expecting to major in art history, the other in writing or psychology. Which, of course, is the whole point of this story, that DNA technology has advanced so far that everyday nonscientists, people like you and the kids down the street, can use it. Our future, indeed
©Txtwriter Inc.