Sequencing DNA: The Sanger Method
Most DNA sequencing is currently done using the “chain termination” technique developed initially by Frederick Sanger, for which he earned his second Nobel Prize (figure 12). (1) A short single-stranded primer is added to the end of a single-stranded DNA fragment of unknown sequence. The primer provides a 3´ end for DNA polymerase. (2) The primed fragment is added, along with DNA polymerase and a supply of all four deoxynucleotides (d-nucleotides), to four synthesis tubes. Each contains a different dideoxynucleotide (dd-nucleotide); such nucleotides lack both the 2´ and the 3´ OH groups and are thus chain-terminating. The first tube, for example, contains ddATP and stops synthesis whenever ddA is incorporated into DNA instead of dATP. Because of the relatively low concentration of ddATP compared to dATP, ddA will not necessarily be added to the first A site; this tube will contain a series of fragments of different lengths, corresponding to the different distances the polymerase traveled from the primer before a ddA was incorporated. (3) These fragments can be separated according to size by electrophoresis. (4) A radioactive label (here dATP*) allows the fragments to be visualized on X-ray film, and the newly made sequence can be read directly from the film. Try it. (5) The original fragment has the complementary sequence.
Techniques such as Southern blotting and PCR enable investigators to identify specific genes and produce them in large quantities, while RFLP analysis and the Sanger method identify individuals and unknown gene sequences. 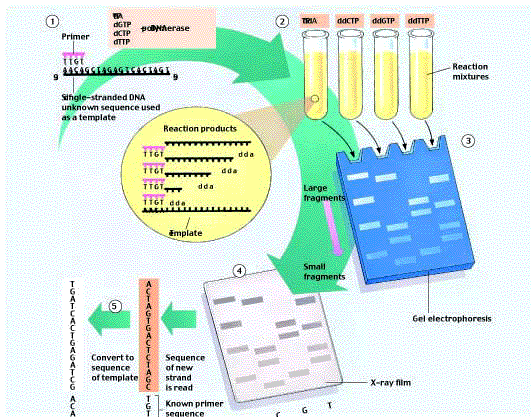 Figure 12
Figure 12
The Sanger dideoxynucleotide sequencing method.