Gene Microarrays
Microarrays
A gene microarray is a square of glass smaller than a postage stamp, covered with millions of strands of DNA arrayed like blades of grass. Microarrays were invented in 1991 by gene scientist Stephen Fodor. In a flash of insight, he saw that photolithography, the process used to etch semiconductor circuits into silicon, could also be used to assemble particular DNA molecules on a chipa gene microarray.
Think of the chip surface as a field of assembly sites, much as a TV screen is a field of colored dots. Just as a scanning beam moves over each individual TV dot instructing it to be red, green, or blue (the three components of color), so a scanning beam moves over each biochip spot, commanding the addition there of a base to growing strands of DNA. A computer using a scanning beam of light to unmask all the sites where an A is to be added, and adds that nucleotide to thousands of growing DNA strands anchored at each of these spots. It then remasks these spots and unmasks instead all the sites scheduled to receive G, then similarly C and T.
When the entire chip has been scanned, each DNA strand has been lengthened one nucleotide unit. The identity of the added nucleotide is in each case determined by the computer, The computer then repeats the process, layer by layer, until each DNA strand is some 25 bases long, enough to unambiguously identify one specific gene. One gene microarray chip made in this way can contain hundreds of thousands of specific gene recognition sequences.
How could you use such a microarray chip to delve into a persons genes? All you would have to do is to obtain a little of the persons DNA, say from a blood sample. Flush fluid containing the DNA over the chip surface. Every place that the DNA has a gene matching one of the microarray strands, it will stick to it in a way the computer can detect.
Patterns of gene expression can be directly determined with gene microarrays. To do this, mRNA isolated from the cells being studied is reverse transcribed to make copy DNA (cDNA). Because the reaction mixture contains fluorescently-labeled nucleotides, any DNA the cDNA binds to is easily recognized. When this labeled cDNA is mixed with a gene microarray representing many thousands of genes, spots light up corresponding to those genes being transcribed in the cells (figure 8).
Researchers are busily comparing the reference sequence of the human genome to the DNA of individual people, and noting any differences they detect. In this way, they are finding SNPs, which we discussed earlier. These SNPs, or spot differences in the identity of particular nucleotides, record every way in which a particular individual differs from the reference sequence. Some SNPs cause diseases like cystic fibrosis or sickle cell anemia. Others may give you red hair or elevated cholesterol in your blood. The human genome tells us that SNPs can be expected to occur at a frequency of about one per thousand nucleotides, scattered about randomly over the chromosomes. Each of us thus differs from the standard “type sequence” in some thirty thousand nucleotide SNPs.
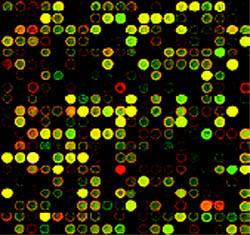
Figure 8 A gene microarray designed to screen for Pax6 activity.
This particular microarray compares gene expression patterns in a fruitfly in response to DNA of the development-regulating gene Pax6 isolated from a fruitfly (labeled with a green fluorescent dye) and from a mouse Pax6 (labeled with a red fluorescent dye). The amount of fluorescence correlates with the amount of gene expression. The yellow spots indicate that these genes were activated in response to both of the Pax6 genes.
How Microarray Gene Chips Can Be Used to Screen for Cancer
One of the biggest decisions facing an oncologist (cancer doctor) treating a tumor is to select the proper treatment. Most cancer cells look alike, although the tumors may in fact be caused by quite different forms of cancer. If the oncologist could clearly identify the cancer, very targeted therapies might be possible. Unable to tell the difference for sure, however, oncologists take no chances. Tumors are treated with therapy that attacks all cancers, usually with severe side effects.
Researchers Todd Golub and Eric Lander took a vital step towards treating cancer, using gene microarrays to sniff out the differences between different forms of a deadly cancer of the immune system.
The ideal way to tell the difference between two kinds of cancer is to compare the mutations that led to the cancer in the first place. The mutations that cause many lung cancers are caused by a tobacco-induced alteration of a single DNA nucleotide in one gene. Such spot differences between the version of a gene one person has and another person has, or a cancer patient has, are examples of medically important SNPs. Golub and Lander obtained bone marrow cells from patients with two types of leukemia (cancer of white blood cells), and exposed DNA from each to microarrays containing 6,817 human genes. Using high-speed computer programs, Golub and Lander examined each of the 6,817 positions on the chip (continued on next page).