Last week Boston researchers Todd Golub and Eric Lander took a vital step towards treating cancer, using new DNA technology to sniff out the differences between different forms of cancer.
One of the biggest decisions facing an oncologist (cancer doctor) treating a tumor is to select the proper treatment. Most cancer cells look alike, although the tumors may in fact be caused by quite different forms of cancer. If the oncologist could clearly identify the cancer, very targeted therapies might be possible. Unable to tell the difference for sure, however, oncologists take no chances. Tumors are treated with therapy that attacks all cancers, usually with severe side effects.
Golub and Lander worked with little glass microchips the size of postage stamps. Etched on the glass are lines. Within each line are layers of DNA, each a gene or a part of a gene. Every known gene in the human body lies somewhere on the DNA microchip, 6,817 in all.
The way to tell the difference between two kinds of cancer is to compare the gene changes that led to the cancer in the first place. Biologists call such gene changes mutations. The mutations that cause many lung cancers are caused by tobacco-induced alteration of a single DNA nucleotide (the genetic message in your genes is written in code letters called nucleotides.) in one gene. Such spot differences between the version of a gene I have and the one you do, or a cancer patient does, are called “single nucleotide polymorphisms,” or SNPs.
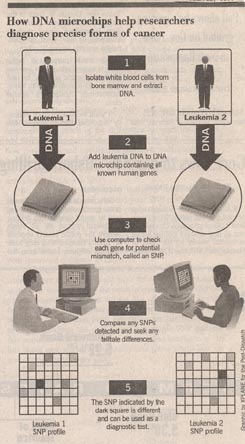
Golub and Lander obtained bone marrow cells from patients with two types of leukemia (cancer of white blood cells), and exposed DNA from each to the DNA microchip. The DNA in the chip interacts in different ways with a sample of DNA placed upon the chip, depending wether the sample and chip DNAs are the same or not. Using high-speed computer programs, Golub and Lander examined each of the 6,817 positions on the chip. The two forms of leukemia each showed gene changes from normal, but, importantly, the changes were different in each case! Each had their own characteristic SNP.
The DNA microchip thus offers a quick and reliable way to identify any type of cancer. Just look and see what SNP is present.
Gene chip technology is likely to dominate medicine in the coming millennium, a prospect both exciting and scary. As the human genome project charges toward completion within the next few years, its researchers are excitedly assembling a huge database of SNPs, single-character typos in the complement of human genes we all carry. Research indicates that SNPs can be expected to occur at a frequency of about one per thousand nucleotides, scattered about randomly over the chromosomes. Each of us thus differs from the standard “type sequence” in several thousand nucleotide SNPs.
Researchers have announced plans to compile a data base of hundreds of thousands of SNPs over the next two years. Screening SNPs and comparing them to known SNP data bases will soon allow doctors to screen each of us for copies of genes leading to genetic diseases. Many genetic diseases are associated with SNPs, including cystic fibrosis and muscular dystrophy.
Most of the few thousand SNPs we each carry do no damage, cause no disease. But if you think about it, they are responsible for much of the difference between individuals. My SNPs are the part of me that makes me ME. Studies of how personality and intelligence (however measured) are inherited among humans indicate that over three quarters of the variation among individuals is due to heredity, to differences in the DNA — to SNPs.
So what does it mean to say that soon someone will be able, on a DNA microchip, to read your SNP profile? Sitting at a computer terminal, someone you have never met may inspect each SNP and ask if it matches anything in the data base. Is it correlated with any disease, for example, or with any tendency to excessive weight or other medically significant condition? Do you really want to know? Do you want your insurance company to know?
Any even more troublesome scenario will become possible as more and more people have their SNP profiles added to the data base. Many of the most interesting things that our parents contribute to what we are like are complex, hereditary gifts that involve many genes, not single SNPs. But with a big enough data base, genomic scientists will be able to begin searching the data base for multiple-SNP patterns that correlate with personality, behavior, and intelligence. In this brave new world, individuality and privacy will take on a whole new meaning.
©Txtwriter Inc.